HDF5 in LabVIEW

Yannic Risters
In one way or another, every LabVIEW application deals with data. This includes the acquisition, display, processing, and storage of data. Depending on the application, it may be necessary to manage a huge amount of data. It may also be required to store the data as fast as possible, as efficiently as possible and in a way that it can be used by different programs for example MATLAB or Python for further data analysis.
So choosing the appropriate file format is crucial. There are different file formats to choose from like ASCII, Binary, XML, and TDMS. NI provides a comparison for these file formats:
https://www.ni.com/nl-nl/innovations/white-papers/09/comparing-common-file-i-o-and-data-storage-approaches.html
According to this comparison, all of these file formats have their pros and cons. In general, TDMS appears to address the shortcomings of the other file formats.
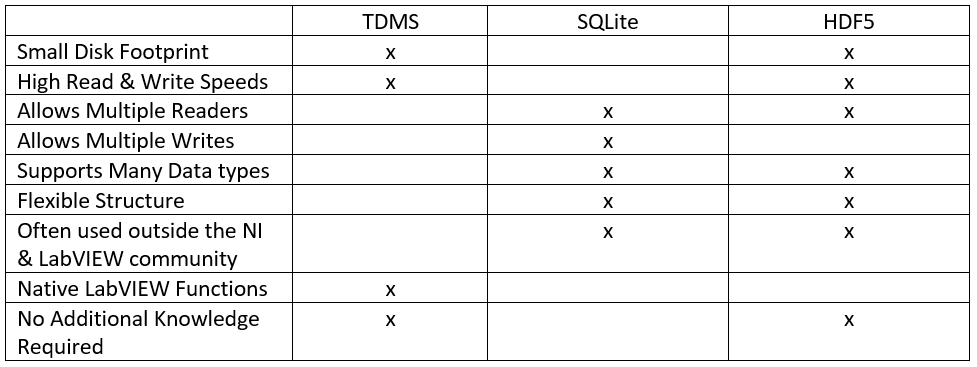
If it comes to large amounts of data, TDMS is indeed one possible option. LabVIEW includes native TDMS functions that are relatively easy to use and without additional knowledge. It also has a small disk footprint, and high reading and writing speeds. However, TDMS does not allow multiple readers and writers at the same time. Furthermore, only a limited number of data types can be written to a TDMS file, so the file structure based on groups and channels has limited flexibility. Beyond that, TDMS is not often used outside the NI and LabVIEW communities.
Another possibility could be the use of SQLite and databases. SQLite allows multiple readers and writers, it supports multiple data types, the structure of e.g. tables is rather flexible, and it is often used outside the NI and LabVIEW community. However, SQLite does not necessarily have a small disk footprint, high reading and writing speeds. Furthermore, LabVIEW does not include native SQLite functions and it is necessary to install SQLite toolkits for LabVIEW. Beyond that, SQLite requires additional knowledge about how to write queries.
A third option considering large amounts of data could be the use of HDF5. To some extent, this file format is comparable to TDMS. However, one major difference is that the file structure of HDF5 is relatively flexible e.g. considering the usage of groups and subgroups. In addition, HDF5 allows many data types, multiple readers at the same time, and it is often used outside the NI and LabVIEW community. Furthermore, no additional knowledge is required to use HDF5 functions. But LabVIEW does not include native HDF5 functions and it is required to install HDF5 toolkits for LabVIEW.
In this blog post, I want to give you an insight into HDF5 and how to use it in LabVIEW.
So, what is HDF5?
The Hierarchical Data Format (HDF) is a group of file formats designed to store and organize large amounts of data. Its development started in 1987 when the Graphics Foundations Task Force (GFTF) at the National Center for Supercomputing Applications (NCSA) was looking for a new data format for scientific purposes. As time went by, the data format further evolved, resulting in the development of HDF5, the most recent HDF version.
The HDF file format is maintained by the HDF Group, a non-profit company that works on the further development of HDF technology, and that ensures long-term and continued access to data, stored in HDF.
https://www.hdfgroup.org/
HDF5 has some interesting characteristics, among which:
- The data is stored in binary format which saves storage space.
- There are no limitations considering the number or size of data objects. This means that it can be used to store very large amounts of data.
- HDF5 is high-performing considering I/O.
- It supports dimensional datasets.
- HDF5 allows to include metadata with the data which makes it self-describing data.
- It can be easily shared since, among others, both the data and metadata are included in one file.
- HDF5 is open-source.
- It is cross-platform, and it is supported by, among others, C, C++, MATLAB, and Python.
What is the HDF5 file structure?
HDF5 files are hierarchically structured, and they consist of three primary structures: Groups, datasets, and attributes.
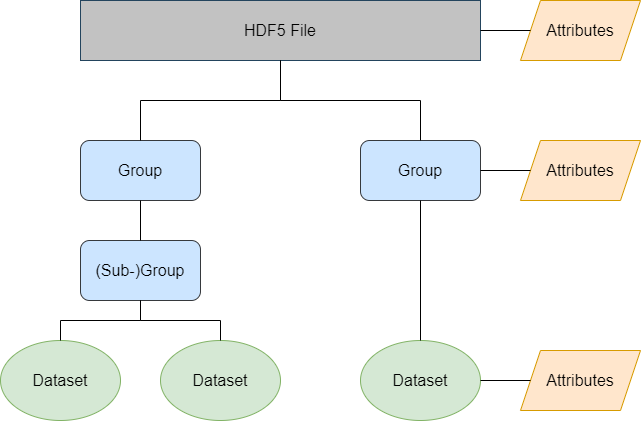
A group is a data object that includes zero or more HDF objects, like subgroups and datasets. It consists of a:
- Header (i.e. the group name and a list of group attributes)
- Symbol table (i.e. a list of the HDF objects belonging to that group)
A dataset is a collection of data, and it consists of a:
- Header:
- Dataset Name
- Datatype (e.g. Integers, Floating point, Strings)
- Dataspace (i.e. the dimensionality of the dataset)
- Storage Layout (Compact or Chunked)
- Dataset attributes
- Data array
An attribute is a small dataset that provides metadata and that can be attached to an HDF5 file, a group, or a dataset. It consists of a:
How to view the content of an HDF5 file?
As previously mentioned, HDF data is stored in binary format which saves storage space. This means, that you cannot view the data in e.g. a simple text editor.
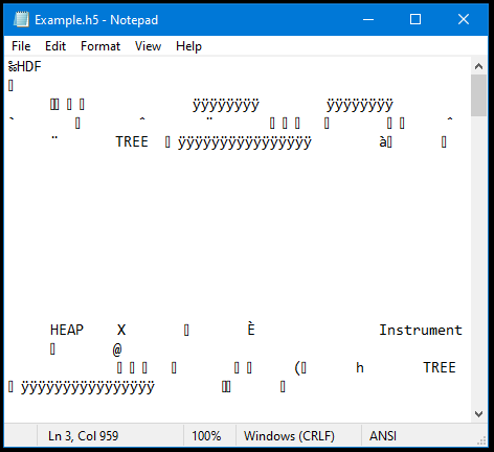
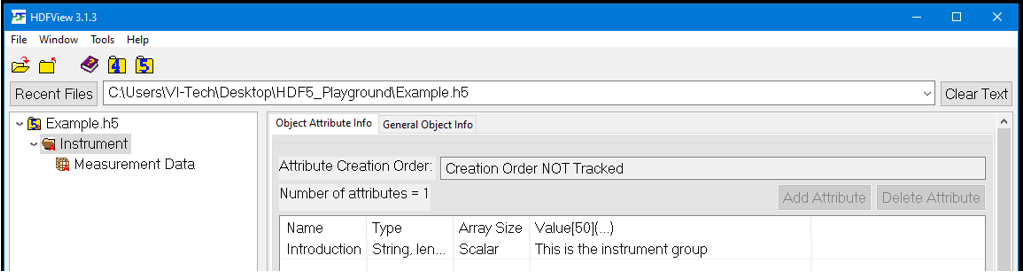
One way to view HDF data is by using the HDF View tool provided by the HDF Group.
https://www.hdfgroup.org/downloads/hdfview/
How to use HDF5 in LabVIEW?
At this moment, there are some LabVIEW toolkits available that could be used for this purpose. One of these toolkits is the “h5labview” package developed by Martijn Jasperse.
https://h5labview.sourceforge.io/?home
It is an open-source toolkit that provides a compact library of functions for using HDF data in LabVIEW.
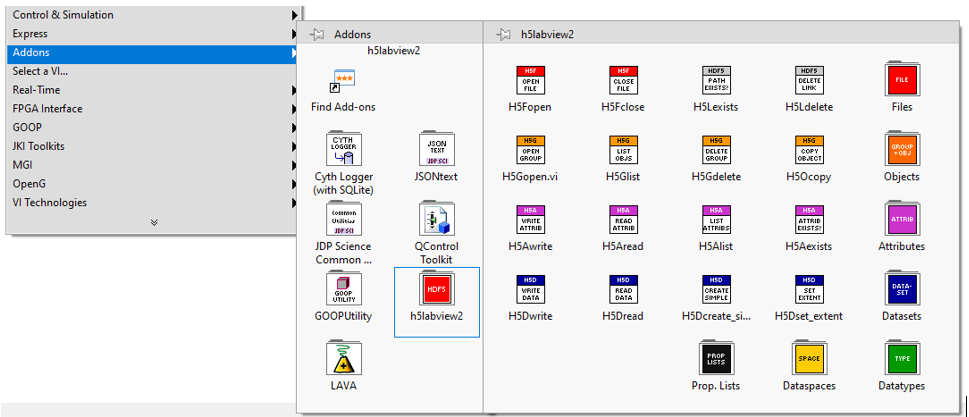
This toolkit is available for LabVIEW 2010 and newer, and it is available for 32-and 64-bit versions of Windows and Linux. From version 2 onwards, h5labview supports several data types including compound data types (e.g. arrays of clusters).
This sounds very nice. But how to actually use this toolkit?
Suppose that you want to measure the temperature and relative humidity (RH) in your office room every hour. For this purpose, you want to use a Raspberry Pi and an I2C sensor. You have already developed an application that reads these quantities using the Raspberry Pi and an I2C sensor, and you now want to extend the application by writing the data to an HDf5 file.
The measurement data has the following structure:
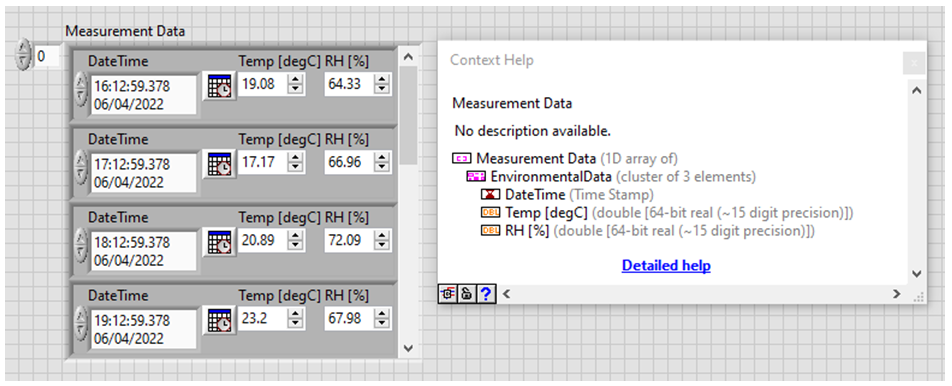
And the HDF5 file should be structured as followed:
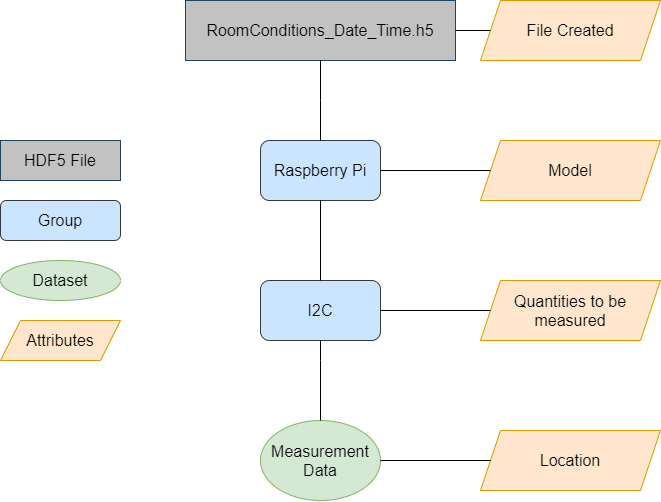
In short, one possible way of using the h5labview toolkit for this use case looks as follows:
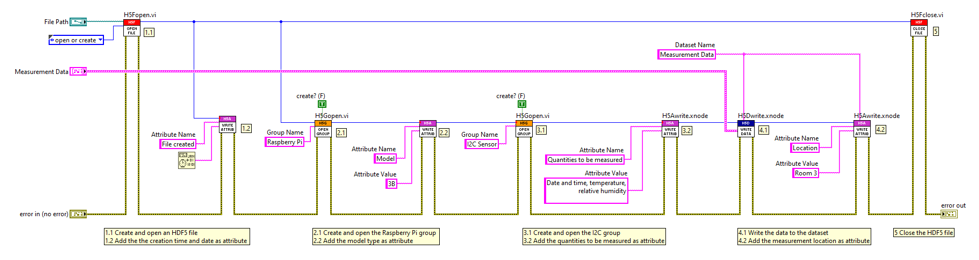
Download the example here: HDF5_LabVIEW_Example.zip
After running the application, you may inspect the resulting HDF5 file. First, you may check the attributes of the file itself:

Second, you may check the attributes of the “Raspberry Pi” group.
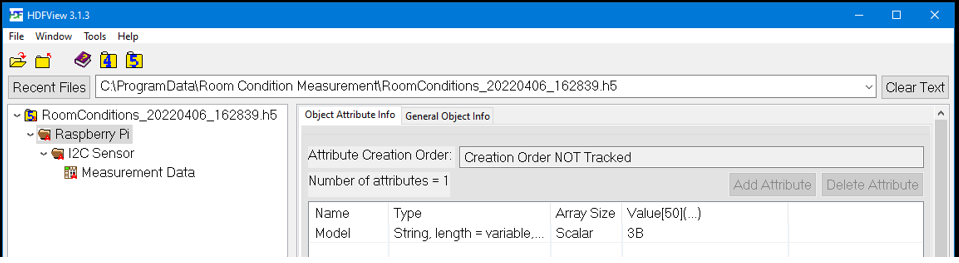
Third, you may check the attributes of the “I2C” group.
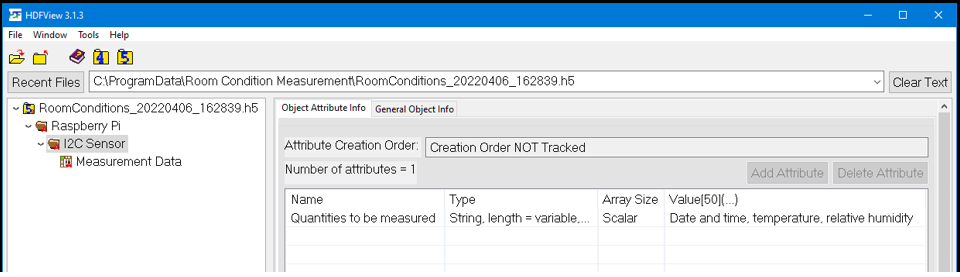
And last, you may check the attributes and data of the dataset called “Measurement data”.
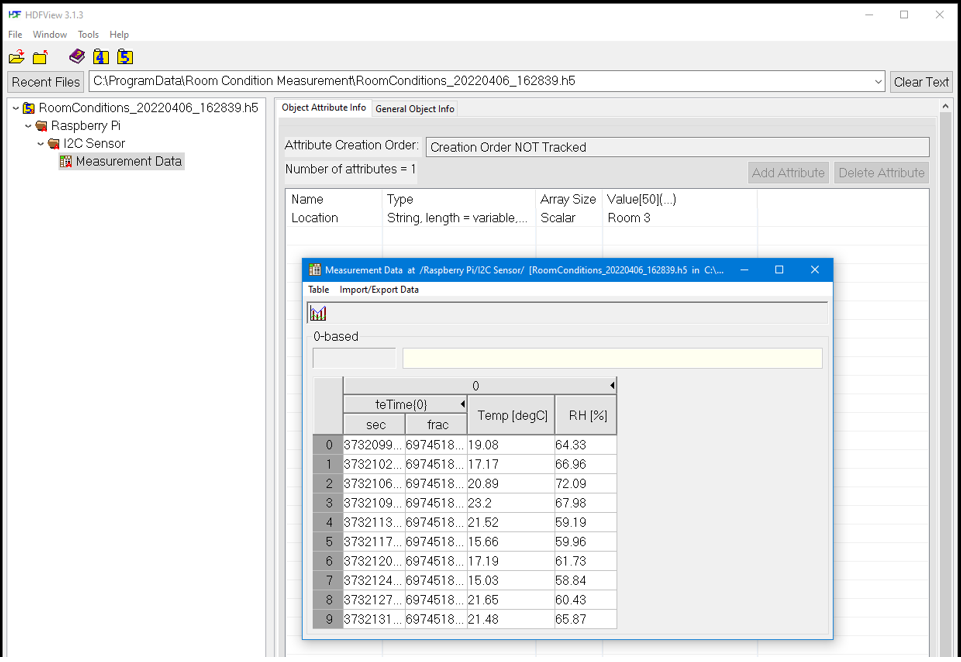
Finally
I hope that you now got an impression about HDF5 and how to use this file format in LabVIEW. You can try it out the next time that you have to work with large amounts of data.

